2026-05-04 ペンシルベニア州立大学(Penn State)
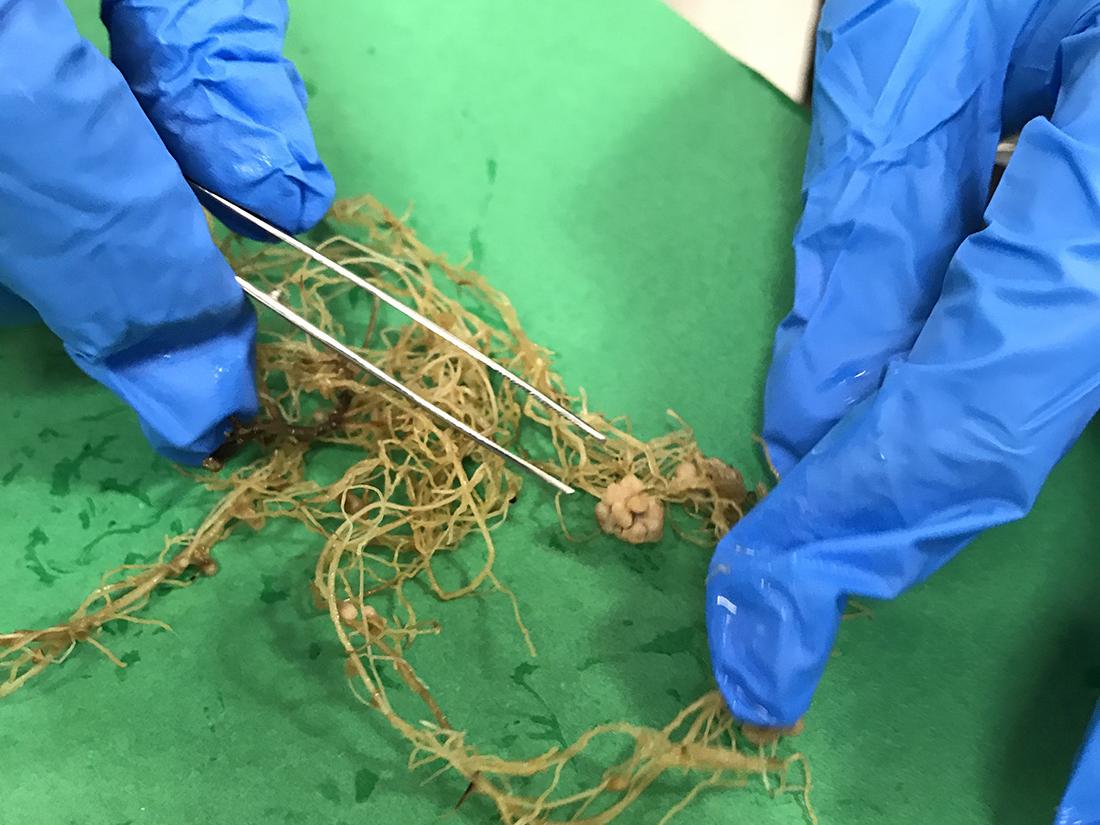
Nodules in legume roots are shown. Rhizobia live inside these specialized structures where they convert atmospheric nitrogen into a form the plant can use. Credit: Penn State. Creative Commons
<関連情報>
- https://www.psu.edu/news/research/story/plant-genes-influence-bacterial-evolution-legume-bacteria-partnership
- https://academic.oup.com/ismej/article/20/1/wrag005/8466334?login=false
共生に影響を与えるマメ科植物の遺伝子の変異は、根粒菌共生体にとって複雑な選択環境を作り出す Mutations in legume genes that influence symbiosis create a complex selective landscape for rhizobial symbionts
Sohini Guha,Regina B Bledsoe,Jeremy Sutherland,Brendan Epstein,Gwendolyn M Fry,Vikram Venugopal,Siva Sankari,Alejandra Gil-Polo,Garrett Levin,Barney A Geddes,…
The ISME Journal Published::06 February 2026
DOI:https://doi.org/10.1093/ismejo/wrag005
Abstract
In the mutualism between leguminous plants and rhizobial bacteria, rhizobia live inside root nodules, creating potential for host genes to shape the rhizobial selective environment. Many host genes that affect symbiosis have been identified; however, the extent to which these genes affect selection acting on rhizobia is unknown. In this study, we inoculated 18 Medicago truncatula symbiotic mutants (including mutants that alter Nodule Cysteine-Rich (NCR) peptide production, plant defence, and nodule number regulation) with a mixture of 86 Sinorhizobium meliloti strains. Most mutations resulted in reduced host benefits, but the effects on rhizobial benefit (i.e. relative strain fitness) varied widely, revealing widespread host-by-strain fitness interactions. Genome-wide association analyses identified variants on rhizobial replicons pSymA and pSymB as important in mediating strain fitness responses to host mutations. Whereas most top variants affected rhizobial fitness with one host mutation (limited effect variants), nine affected fitness across six or more host mutations. These pervasive variants occurred primarily on pSymA, the symbiotic replicon, and include fixL and some metabolic genes. In contrast to the limited effect variants, variants with pervasive positive effects on strain fitness when host genes were mutated tended to adversely affect fitness in wild-type hosts. Competition assays across Medicago genotypes confirmed a pervasive role for one candidate (malonyl-CoA synthase), and AlphaFold multimer modelling suggests that many rhizobial top candidates could interact with host NCR peptides. Our results reveal how host genetic mutations alter strain fitness, setting the stage for improving rhizobial inoculants and breeding legume hosts better adapted to multi-strain environments.


