2026-03-17 名古屋大学
<関連情報>
- https://www.nagoya-u.ac.jp/researchinfo/result/2026/03/post-962.html
- https://www.nagoya-u.ac.jp/researchinfo/result/upload_images/20260317_agr.pdf
- https://www.pnas.org/doi/10.1073/pnas.2527570123
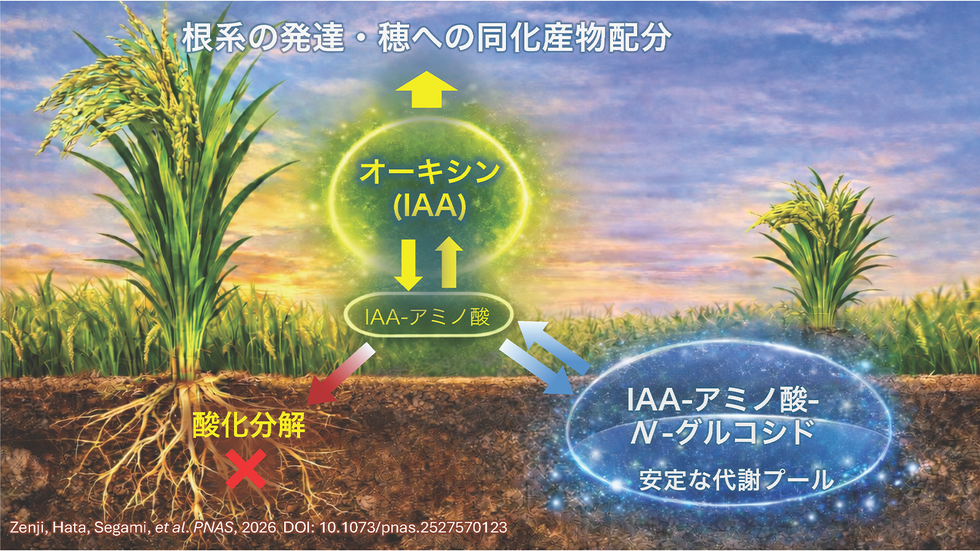
インドール-3-アセチルアミノ酸のN-グルコシル化は、イネ(Oryza sativa)のオーキシン代謝と成長特性を調節する N-glucosylation of indole-3-acetyl amino acids modulates auxin metabolism and growth traits in Oryza sativa
Anna Zenji, Kimihiko Hata, Shoji Segami, +12 , and Hitoshi Sakakibara
Proceedings of the National Academy of Sciences Publishe:March 18, 2026
DOI:https://doi.org/10.1073/pnas.2527570123
Significance
Although auxin metabolism is central to the regulation of plant growth and development, the fate of conjugates beyond oxidation has remained unclear. Here, we identify a rice UDP-glucosyltransferase, IAAspGT, that catalyzes the N-glucosylation of indole-3-acetyl amino acid conjugates. This modification diverts auxin conjugates from irreversible oxidation, establishing a secondary regulatory pool that fine-tunes auxin availability. Natural allelic variation in IAAspGT explains the differences in root system growth, coleoptile elongation, and biomass allocation between a japonica– and an indica-type cultivar. The ancestral high-activity allele underscores the evolutionary significance and agricultural potential of this overlooked pathway, offering prospects for crop improvement.
Abstract
Dynamic regulation of auxin (indole-3-acetic acid; IAA) levels is crucial for proper plant growth and development and is finely regulated through biosynthesis, transport, metabolic inactivation, and signal transduction. While O-glucosylation of IAA is well established, the physiological significance of N-glucosylation pathways acting on IAA-amino acid conjugates remains largely unexplored. Here, we identify a UDP-glucosyltransferase in rice (Oryza sativa), designated IAAspGT, that catalyzes the N-glucosylation of indole-3-acetyl (IA) amino acid conjugates. Functional analysis revealed that natural allelic variants of IAAspGT differ markedly in enzymatic activity, with the high-activity allele prevalent in indica and the low-activity allele in japonica cultivars. Mutational analysis identified key residues within the glucose-accepting substrate-binding domain that account for this variation. A metabolic perturbation experiment demonstrated that IAAspGT competes with the DAO-mediated oxidation, thereby affecting active auxin levels. Plants harboring the high-activity allele exhibited enhanced root elongation under nutrient-deficient conditions and altered growth allocation between the panicle and other above-ground organs. Haplotype analysis indicated that the high-activity allele is ancestral and widely retained in the wild relatives. These findings establish IA-amino acid N-glucosylation as a previously overlooked metabolic branch that modulates auxin availability and contributes to environmental responses and growth plasticity in rice.